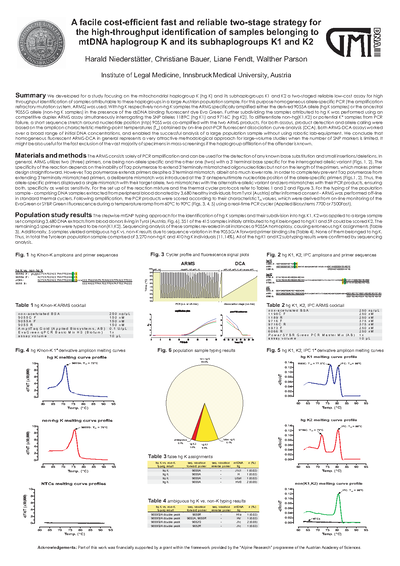
A facile cost-efficient fast and reliable two-stage strategy for the high-throughput identification of samples belonging to mtDNA haplogroup K and its subhaplogroups K1 and K2
Niederstätter,H.; Bauer,C.; Fendt,L.,Parson,W.
We developed for a study focusing on the mitochondrial haplogroup K (hg K) and its subhaplogroups K1 and K2 a two-staged reliable low-cost assay for high throughput identification of samples attributable to these haplogroups in a large Austrian population sample. For this purpose homogeneous allele-specific PCR (the amplification refractory mutation system, ARMS) was used. With hg K respectively non-hg K samples the ARMS specifically amplified either the derived 9055A allele (hg K samples) or the ancestral 9055G allele (non-hg K samples) in the presence of the dsDNA binding fluorescent dye Eva Green. Further subdividing the samples attributed to hg K was performed using an competitive duplex ARMS assay simultaneously interrogating the SNP alleles 1189C (hg K1) and 9716C (hg K2). To differentiate non-hg(K1,K2) or potential K* samples from PCR failure, a short sequence stretch around nucleotide position (ntp) 9055 was co-amplified with the two ARMS products. For both assays, product detection and allele calling were based on the amplicon characteristic melting-point temperatures (T m ) obtained by on-line post-PCR fluorescent dissociation curve analysis (DCA). Both ARMS-DCA assays worked over a broad range of initial DNA concentrations, and enabled the successful analysis of a large population sample without using robotic lab-equipment. We conclude that homogeneous fluorescent ARMS-DCA in general represents a very attractive methodological approach for large-volume studies when the number of SNP markers is limited. It might be also useful for the fast exclusion of the vast majority of specimens in mass-screenings if the haplogroup affiliation of the offender is known.